AR1 Transformer Function
AR1_transform.RdThe transformer function takes a bamlss.frame object and transforms
the response and the design matrices to account for lag 1 autocorrelation. The method
is also known as Prais-Winsten estimation.
trans_AR1(rho = 0.1)
AR1(rho = 0.1)Arguments
- rho
Specifies the correlation parameter at lag 1.
Value
A transformer function which can be used in the bamlss call.
References
Johnston, John (1972). Econometric Methods (2nd ed.). New York: McGraw-Hill. pp. 259--265.
See also
Examples
if (FALSE) ## Simulate AR1 data.
set.seed(111)
n <- 240
d <- data.frame("t" = 1:n)
## Nonlinear function.
f <- function(x) {
2 + sin(x / n * 2 * pi - pi)
}
## Correlated errors.
rho <- 0.8
e <- rnorm(n, sd = 0.1)
u <- c(e[1], rep(NA, n - 1))
for(i in 2:n){
u[i] <- rho * u[i - 1] + e[i]
}
## Response.
d$y <- f(d$t) + u
## Plot time-series data.
plot(d, type = "l")
## Estimate models without and with AR1 transformation.
b0 <- bamlss(y ~ s(t,k=20), data = d, criterion = "BIC")
#> BIC 136.4084 logPost -97.6646 logLik -44.2986 edf 8.7236 eps 0.3974 iteration 1
#> BIC -41.9715 logPost -8.4746 logLik 44.8913 edf 8.7236 eps 0.1812 iteration 2
#> BIC -152.399 logPost 46.7393 logLik 100.1053 edf 8.7236 eps 0.1157 iteration 3
#> BIC -189.094 logPost 75.9567 logLik 119.8168 edf 9.2214 eps 0.0649 iteration 4
#> BIC -221.762 logPost 91.1354 logLik 153.2188 edf 15.449 eps 0.0523 iteration 5
#> BIC -228.519 logPost 93.8649 logLik 162.6250 edf 17.649 eps 0.0209 iteration 6
#> BIC -228.766 logPost 94.0024 logLik 163.9364 edf 18.083 eps 0.0033 iteration 7
#> BIC -228.770 logPost 94.0090 logLik 164.1063 edf 18.144 eps 0.0002 iteration 8
#> BIC -228.770 logPost 94.0093 logLik 164.1187 edf 18.148 eps 0.0000 iteration 9
#> BIC -228.770 logPost 94.0093 logLik 164.1187 edf 18.148 eps 0.0000 iteration 9
#> elapsed time: 0.08sec
#> Starting the sampler...
#>
#> | | 0% 1.19sec
#> |* | 5% 3.38sec 0.18sec
#> |** | 10% 2.24sec 0.25sec
#> |*** | 15% 1.88sec 0.33sec
#> |**** | 20% 1.62sec 0.40sec
#> |***** | 25% 1.43sec 0.48sec
#> |****** | 30% 1.33sec 0.57sec
#> |******* | 35% 1.19sec 0.64sec
#> |******** | 40% 1.07sec 0.71sec
#> |********* | 45% 0.95sec 0.78sec
#> |********** | 50% 0.85sec 0.85sec
#> |*********** | 55% 0.75sec 0.92sec
#> |************ | 60% 0.65sec 0.98sec
#> |************* | 65% 0.57sec 1.05sec
#> |************** | 70% 0.48sec 1.11sec
#> |*************** | 75% 0.40sec 1.19sec
#> |**************** | 80% 0.32sec 1.27sec
#> |***************** | 85% 0.24sec 1.35sec
#> |****************** | 90% 0.16sec 1.42sec
#> |******************* | 95% 0.08sec 1.50sec
#> |********************| 100% 0.00sec 1.59sec
b1 <- bamlss(y ~ s(t,k=20), data = d, criterion = "BIC",
transform = AR1(rho = 0.8))
#> BIC 120.6978 logPost -100.564 logLik -19.7587 edf 14.812 eps 0.4482 iteration 1
#> BIC -183.664 logPost 54.4906 logLik 112.0547 edf 7.3797 eps 0.1809 iteration 2
#> BIC -316.134 logPost 120.7259 logLik 178.2900 edf 7.3797 eps 0.1060 iteration 3
#> BIC -372.549 logPost 148.9333 logLik 206.4974 edf 7.3797 eps 0.0648 iteration 4
#> BIC -381.197 logPost 153.2571 logLik 210.8212 edf 7.3797 eps 0.0266 iteration 5
#> BIC -380.659 logPost 169.9073 logLik 211.2022 edf 7.6169 eps 0.0059 iteration 6
#> BIC -380.659 logPost 169.9076 logLik 211.2025 edf 7.6169 eps 0.0002 iteration 7
#> BIC -380.659 logPost 169.9076 logLik 211.2025 edf 7.6169 eps 0.0000 iteration 8
#> BIC -380.659 logPost 169.9076 logLik 211.2025 edf 7.6169 eps 0.0000 iteration 8
#> elapsed time: 0.10sec
#> Starting the sampler...
#>
#> | | 0% 1.19sec
#> |* | 5% 1.22sec 0.06sec
#> |** | 10% 1.15sec 0.13sec
#> |*** | 15% 1.07sec 0.19sec
#> |**** | 20% 1.03sec 0.26sec
#> |***** | 25% 1.03sec 0.34sec
#> |****** | 30% 1.21sec 0.52sec
#> |******* | 35% 1.09sec 0.58sec
#> |******** | 40% 1.00sec 0.67sec
#> |********* | 45% 0.90sec 0.73sec
#> |********** | 50% 0.80sec 0.80sec
#> |*********** | 55% 0.71sec 0.87sec
#> |************ | 60% 0.62sec 0.94sec
#> |************* | 65% 0.54sec 1.01sec
#> |************** | 70% 0.47sec 1.11sec
#> |*************** | 75% 0.39sec 1.18sec
#> |**************** | 80% 0.31sec 1.24sec
#> |***************** | 85% 0.23sec 1.31sec
#> |****************** | 90% 0.15sec 1.38sec
#> |******************* | 95% 0.08sec 1.46sec
#> |********************| 100% 0.00sec 1.55sec
## Estimate full AR1 model.
b2 <- bamlss(y ~ s(t,k=20), data = d, criterion = "BIC",
family = "AR1")
#> BIC 128.8853 logPost -98.9894 logLik -37.7967 edf 9.7236 eps 0.5982 iteration 1
#> BIC -80.2651 logPost 5.5858 logLik 66.7784 edf 9.7236 eps 0.1685 iteration 2
#> BIC -246.753 logPost 88.8299 logLik 150.0225 edf 9.7236 eps 0.1025 iteration 3
#> BIC -339.747 logPost 135.3267 logLik 196.5194 edf 9.7236 eps 0.0643 iteration 4
#> BIC -363.204 logPost 147.0552 logLik 208.2479 edf 9.7236 eps 0.0326 iteration 5
#> BIC -348.719 logPost 157.7974 logLik 209.1956 edf 12.712 eps 0.0083 iteration 6
#> BIC -348.723 logPost 157.7992 logLik 209.1975 edf 12.712 eps 0.0006 iteration 7
#> BIC -348.723 logPost 157.7993 logLik 209.1975 edf 12.712 eps 0.0001 iteration 8
#> BIC -348.723 logPost 157.7993 logLik 209.1975 edf 12.712 eps 0.0000 iteration 9
#> BIC -348.723 logPost 157.7993 logLik 209.1975 edf 12.712 eps 0.0000 iteration 9
#> elapsed time: 0.16sec
#> Starting the sampler...
#>
#> | | 0% 1.67sec
#> |* | 5% 2.32sec 0.12sec
#> |** | 10% 2.03sec 0.23sec
#> |*** | 15% 1.97sec 0.35sec
#> |**** | 20% 1.82sec 0.45sec
#> |***** | 25% 1.67sec 0.56sec
#> |****** | 30% 1.53sec 0.65sec
#> |******* | 35% 1.40sec 0.75sec
#> |******** | 40% 1.28sec 0.85sec
#> |********* | 45% 1.16sec 0.95sec
#> |********** | 50% 1.05sec 1.05sec
#> |*********** | 55% 0.94sec 1.15sec
#> |************ | 60% 0.84sec 1.26sec
#> |************* | 65% 0.74sec 1.37sec
#> |************** | 70% 0.63sec 1.48sec
#> |*************** | 75% 0.53sec 1.58sec
#> |**************** | 80% 0.42sec 1.67sec
#> |***************** | 85% 0.31sec 1.77sec
#> |****************** | 90% 0.21sec 1.87sec
#> |******************* | 95% 0.10sec 1.97sec
#> |********************| 100% 0.00sec 2.06sec
rho <- predict(b2, model = "rho", type = "parameter")
print(range(rho))
#> [1] 0.7360218 0.7360218
## Estimated standard deviations.
sd0 <- predict(b0, model = "sigma", type = "parameter")
sd1 <- predict(b1, model = "sigma", type = "parameter")
sd2 <- predict(b2, model = "sigma", type = "parameter")
print(round(c(sd0[1], sd1[1], sd2[1]), 2))
#> [1] 0.13 0.00 0.10
## Plot fitted trends.
p0 <- predict(b0, model = "mu")
p1 <- predict(b1, model = "mu")
p2 <- predict(b2, model = "mu")
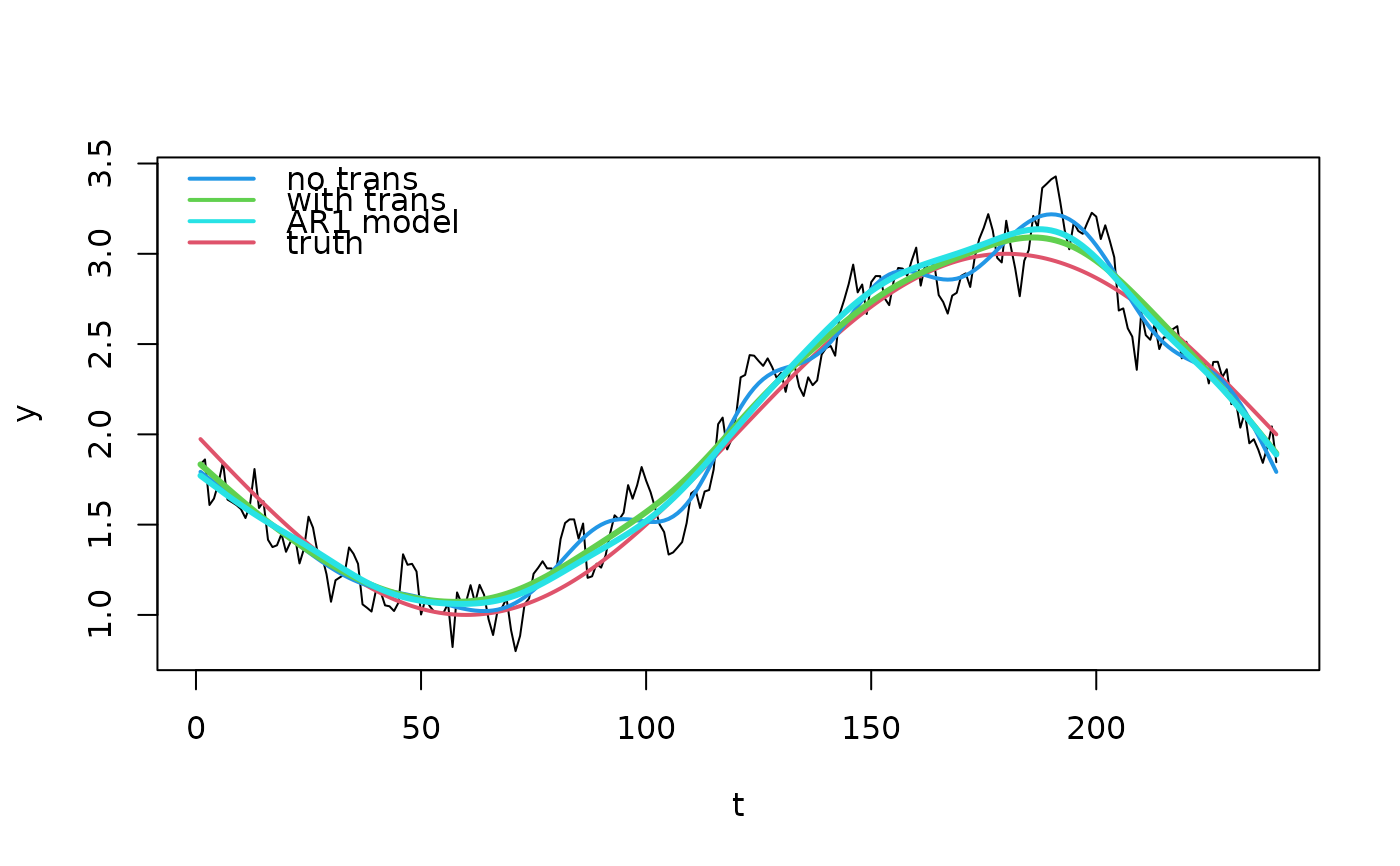
plot(d, type = "l")
lines(f(d$t) ~ d$t, col = 2, lwd = 2)
lines(p0 ~ d$t, col = 4, lwd = 2)
lines(p1 ~ d$t, col = 3, lwd = 3)
lines(p2 ~ d$t, col = 5, lwd = 3)
legend("topleft",
c("no trans", "with trans", "AR1 model", "truth"),
lwd = 2, col = c(4, 3, 5, 2), bty = "n")